Further extending
our use of JSpecView to monitor our reactions by NMR, we have been able to generate good kinetics information for the first step of the Ugi reaction, when an imine forms.
We learned that imine formation is about ten times faster in
methanol than in
chloroform and that our aromatic aldehydes react about 3 orders of magnitude slower than
phenylacetaldehyde. Furthermore,
phenylacetaldehyde suffers from so many side reactions on the time scale of our experiments that we probably won't revisit it until we are comfortable with the well behaved aromatic aldehydes.
Even though we have not yet obtained our
Ugi products, this information certainly could be leveraged for other purposes. In fact there is so much information in these NMR monitoring spectra that it reminds me of the situation in bioinformatics where large volumes of data (e.g. genomic) are generated and shared for unlimited analysis by the larger community.
In a typical organic chemistry paper, the experimental section only reports on the quantity of starting materials, a summary of the reaction conditions, the purification and the yield and characterization of the final products. Additional information is usually omitted if it does not fit into the arguments of the paper deemed essential by the author and reviewers.
In order to be able to understand a reaction it is necessary to be able to dissect and interrogate as much of its information space as possible. A low yield for a particular reaction is very little to go on when it is not possible to test hypotheses of possible side reactions.
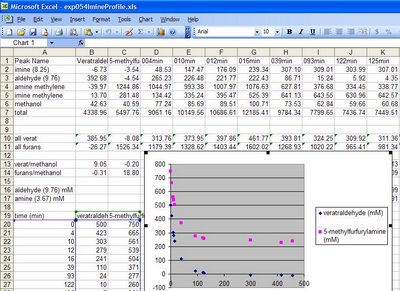
A good example of a surprising (to me at least) insight on our attempts at the Ugi reaction is shown in the image below (from
EXP046). This is a plot of the concentration of imine and aldehyde after addition of the acid and isonitrile. Even though the imine formation was confirmed to be near complete (85%), addition of the last two components actually
reversed the ratio of aldehyde to imine over the following 2 hours, only to be reversed again over the course of several days. The aldehyde concentration did not only change relative to the imine but it actually
increased in absolute terms over the first 2 hours. Furthermore, the combined quantity of imine and aldehyde remained steady after the first 2 hours, indicating that if the Ugi product did form to some extent, it did not continue after this initial brief period.
In order to generate this plot, we had to manually read the integration of the aldehyde and imine peaks and relate those to the integration of the entire spectrum. This is tedious and probably fairly prone to human error. However, since the raw data are openly available, I would hope that any significant errors would get flagged at some point.
For that reason, as well as the amount of data being generated, this would seem to be a good place to seriously interface with automation. We are currently representing the NMR spectra as compressed JCAMP files, human readable using the JSpecView applet. What I am thinking is that we could stack them all in one file using CML. The standalone JSpecView has the ability to convert JCAMP to several formats, including CML, although there is no command line interface to do this yet (Robert Lancashire is working on it). We would add the basic necessary metadata, such as the chemicals that were used and the elapsed time for each spectrum.
These reaction profile files could be accessed automatically by anyone for any purpose. The first task that I would like to see performed is the plot of the integration for a given spectral area relative to the integration of the entire spectrum. For example, in the case of the spectra below, the aldehyde profile would be generated from the 9.0-10.0 ppm range and the imine from the 8.0-9.0 ppm range.
I suspect that in the near future it will be easier for organic chemists to just automate the execution of experiments with rich data outputs rather than walk the minefield of copyrighted materials and commercial databases for a few measly data points.
 Tags1/C6H9NO/c1-5-2-3-6(4-7)8-5/h2-3H,4,7H2,1H35-methylfurfurylamine1/C9H10O3/c1-11-8-4-3-7(6-10)5-9(8)12-2/h3-6H,1-2H3veratraldehyde
Tags1/C6H9NO/c1-5-2-3-6(4-7)8-5/h2-3H,4,7H2,1H35-methylfurfurylamine1/C9H10O3/c1-11-8-4-3-7(6-10)5-9(8)12-2/h3-6H,1-2H3veratraldehyde